
(Sexual Swimmers)
4 Swimmers
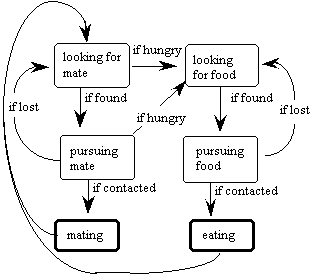
4.1 Perception
A swimmer's field of view covers 360 degrees, and so it can detect other swimmers or food from any direction. There is a limit to how far it can see - its view radius. A swimmer can recognize only the following:• relative orientation of a food bit within its field of view
• relative orientation of a potential mate's head within its field of view
• the proportions of its favorite color in a potential mate
4.2 Mating
Gender difference is not modeled in the swimmer world. Any swimmer can mate with any other swimmer, with the exception of immediate family members (parents, siblings, and offspring) - this is one of the few constraints imposed upon the swimmers in the design of the model, to discourage pockets of stagnant inbreeding.A chosen mate does not need to reciprocate in the mating pursuit for reproduction to happen. The chosen swimmer may have its own agenda, like searching for a mate of its own, or pursuing a food bit. This fact sometimes requires swimmers to pursue indifferent mates for long distances (which adds pressure for evolution of locomotion skill in the population).
There are no reproductive organs modeled in the swimmer world. Swimmers mate with their heads (this is all metaphysical anyway). The act of mating takes place when the heads of the swimmer and its chosen mate reach a distance less than one swimmer segment length (one "swimmer unit"). Mating results in at least one offspring, whose position is initialized directly between the parents, and whose orientation is random. Often, the pursuing swimmer will choose the same mate again in the next time step. In this case, another offspring is born from the same parents. This continues as long as the pursuing swimmer's energy is above the hungry threshold, and the chosen mate has enough energy to contribute its share of offspring energy. The number of consecutive offspring from the same parents typically range from one to four. There is no gestation period in this model.
Every swimmer is a phenotype, represented by a genotype consisting of 17 genes. Each gene is a real number ranging from 0.0 to 1.0, which is mapped to some real, integer, or Boolean value effecting the embryological process of constructing the phenotype. In a birth, the genotypes from both parents are copied and combined using random crossover points to create the offspring genotype. There is a chance of mutation in the offspring genotype, during reproduction. Mutation rate = 0.01.
4.3 Morphology
Swimmers are modeled as 2D figures comprised of interconnected line segments. There is one node (the head) from which a series of limbs emanate. The number of limbs can range from one to four. Each limb is divided into a series of segments, the number of which can range from two to four. All segments are of the same length.The genotype was designed to utilize as few genes as possible, but generating a wide variety of phenotypes, in the spirit of Dawkins' Blind Watchmaker software (89). There are five morphology genes in the genotype. They affect the following phenotypic features:
• number of limbs
• number of segments per limb
• limb "straddle" (angle between consecutive limbs)
• joint angle (base angle per limb joint)
• limb reflection (when "on", this gene reverses all joint angles in alternate limbs)
Figure 2 shows twenty possible (un-evolved) morphologies. In this illustration, there is no autonomous motion affecting these forms so they appear unnaturally symmetrical.
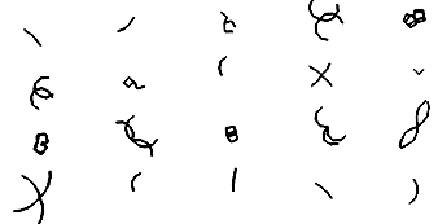
4.4 Motion Control
Each swimmer has an internal motor which drives a series of sinusoidal waves that bend its joints in myriad polyrhythms. In addition to this, the amplitudes and phases of these waves are modulated as a function of the orientation of a chosen food bit or mate, relative to the swimmer (for the purpose of turning). The nine genes dedicated to motion effect the following phenotypic features:• rate (frequency of all wave motions)
• base amplitude for all motions
• limb to limb amplitude offset (adds or subtracts amplitude among consecutive limbs)
• segment to segment amplitude offset (added to the above offset)
• limb to limb phase shift (shifts phases of waves between limbs)
• segment to segment phase shift (added to the above phase shifts)
• turning amplitude (change in amplitude among consecutive limbs for turning)
• turning angle offset (explained below)
• turning phase (change in phase shifts among consecutive limbs for turning)
Each of these are real values which can be negative or positive.
4.5 Reactivity
In pursuit, a swimmer continually compares its axis of orientation to the relative direction of its goal, within its 360 degree visual field. The difference in these directions (an angular value, A) is used to change various parameters in the joint motions from limb to limb. This has the potential effect of causing the swimmer to affect its orientation (whether or not the swimmer rotates so as to move toward its goal depends on many other phenotypic features). Three genes affect a swimmer's ability to turn (the last 3 in the list above). The amount that is added to the amplitudes of all joint motions in a limb for turning is equal to:
limb * cos((A + offset) * pi/180.0) * amp +
(num_limbs-limb+1) * cos((A + offset) * pi/180.0) * amp
where limb ranges from 1 to number_of_limbs, and offset is determined by the gene for limb_turning_angle_offset, and amp is determined by the gene for limb_turning_amplitude. The amount that the phases of waves in a limb's joint motions are shifted for turning is equal to:
limb * cos(A * pi/180.0) * turn_phase
where turn_phase is an angular value, controlled by the gene for limb_turning_phase.
5 Mate Preference
